2.5. Comparative study of BMM method in bivariate linear module using Coleman toy models.#
The best way to learn Taweret is to use it. You can run, modify and experiment with this notebook using GitHub Codespaces.
The models can be found in Coleman Thesis : https://go.exlibris.link/3fVZCfhl
This notebook shows how to use the Bayesian model mixing methods available in bivariate_linear mixing method of package Taweret for a toy problem.
Author : Dan Liyanage
Date : 08/14/2023
import sys
import os
# You will have to change the following imports depending on where you have
# the packages installed
# ! pip install Taweret # if using Colab, uncomment to install
# Setting Taweret path
cwd = os.getcwd()
# Get the first part of this path and append to the sys.path
tw_path = cwd.split("Taweret/")[0] + "Taweret"
sys.path.append(tw_path)
# For plotting
import matplotlib.pyplot as plt
! pip install seaborn # comment if installed
! pip install ptemcee # comment if installed
import seaborn as sns
sns.set_context('poster')
# To define priors. (uncoment if not using default priors)
# ! pip install bilby # uncomment if not already installed
import bilby
# For other operations
import numpy as np
Requirement already satisfied: seaborn in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (0.13.2)
Requirement already satisfied: numpy!=1.24.0,>=1.20 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (2.4.6)
Requirement already satisfied: pandas>=1.2 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (3.0.3)
Requirement already satisfied: matplotlib!=3.6.1,>=3.4 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (3.10.9)
Requirement already satisfied: contourpy>=1.0.1 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (1.3.3)
Requirement already satisfied: cycler>=0.10 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (0.12.1)
Requirement already satisfied: fonttools>=4.22.0 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (4.63.0)
Requirement already satisfied: kiwisolver>=1.3.1 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (1.5.0)
Requirement already satisfied: packaging>=20.0 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (26.2)
Requirement already satisfied: pillow>=8 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (12.2.0)
Requirement already satisfied: pyparsing>=3 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (3.3.2)
Requirement already satisfied: python-dateutil>=2.7 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (2.9.0.post0)
Requirement already satisfied: six>=1.5 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from python-dateutil>=2.7->matplotlib!=3.6.1,>=3.4->seaborn) (1.17.0)
Requirement already satisfied: ptemcee in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (1.0.0)
Requirement already satisfied: numpy in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from ptemcee) (2.4.6)
# Import models with a predict method
from Taweret.models import coleman_models as toy_models
m1 = toy_models.coleman_model_1()
m2 = toy_models.coleman_model_2()
truth = toy_models.coleman_truth()
#!pwd
g = np.linspace(0,9,10)
plot_g = np.linspace(0,9,100)
true_output = truth.evaluate(plot_g)
exp_data = truth.evaluate(g)
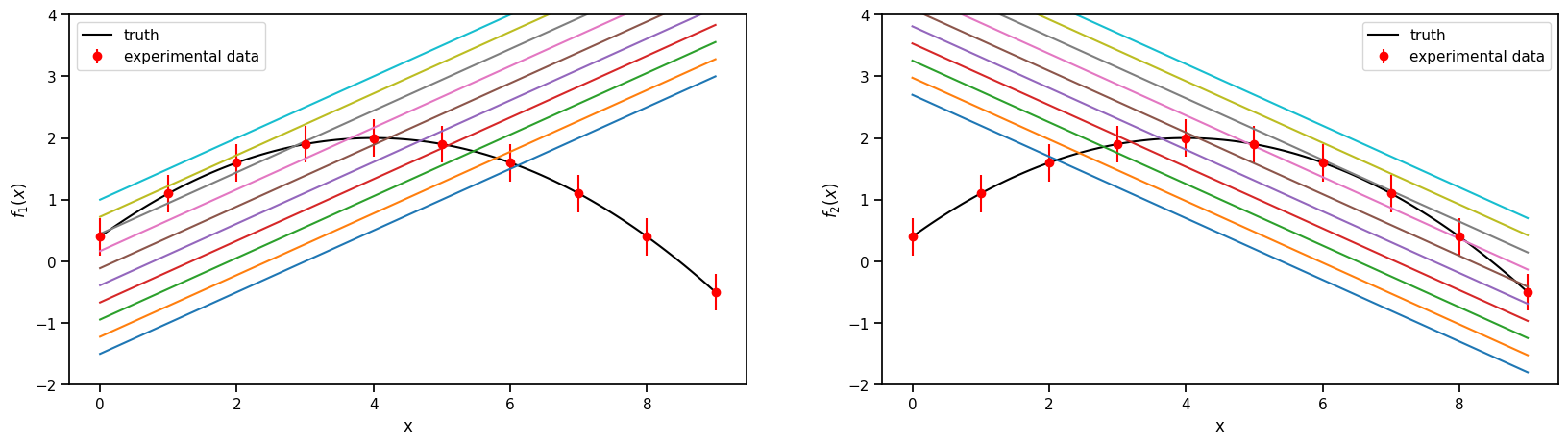
2.5.1. 1. The models and the experimental data.#
Truth
\(f(x) = 2-0.1(x-4)^2\), where \(x \in [-1, 9]\)
Model 1
\(f_1(x,\theta)= 0.5(x+\theta)-2\) , where \(\theta \in [1, 6]\)
Model 2
\(f_2(x,\theta)= -0.5(x-\theta) + 3.7\) , where \(\theta \in [-2, 3]\)
Experimental data
sampled from the Truth with a fixed standard deviation of 0.3
sns.set_context('notebook')
fig, axs = plt.subplots(1,2,figsize=(20,5))
prior_ranges = [(1,6), (-2,3)]
for i in range(0,2):
ax = axs.flatten()[i]
ax.plot(plot_g, true_output[0], label='truth', color='black')
ax.errorbar(g,exp_data[0],exp_data[1], fmt='o', label='experimental data', color='r')
ax.legend()
ax.set_ylim(-2,4)
for value in np.linspace(*prior_ranges[i],10):
if i==0:
predict_1 = m1.evaluate(plot_g, value, full_corr=False)
ax.plot(plot_g, predict_1[0])
ax.set_ylabel(r'$f_1(x)$')
if i==1:
predict_2 = m2.evaluate(plot_g, value, full_corr=False)
ax.plot(plot_g, predict_2[0])
ax.set_ylabel(r'$f_2(x)$')
ax.set_xlabel('x')
2.5.2. 2. Choose a Mixing method#
# Mixing method
from Taweret.mix.bivariate_linear import BivariateLinear as BL
models= {'model1':m1,'model2':m2}
mix_model_BMMC_mix = BL(models_dic=models, method='addstepasym', nargs_model_dic={'model1':1, 'model2':1},
same_parameters = False)
mix_model_BMMcor_mix = BL(models_dic=models, method='addstepasym', nargs_model_dic={'model1':1, 'model2':1},
same_parameters = False, BMMcor=True)
mix_model_mean_mix = BL(models_dic=models, method='addstepasym', nargs_model_dic={'model1':1, 'model2':1},
same_parameters = False, mean_mix=True)
mix_models = [mix_model_BMMC_mix, mix_model_BMMcor_mix, mix_model_mean_mix]
## uncoment to change the prior from the default
priors = bilby.core.prior.PriorDict()
priors['addstepasym_0'] = bilby.core.prior.Uniform(0, 9, name="addstepasym_0")
priors['addstepasym_1'] = bilby.core.prior.Uniform(0, 9, name="addstepasym_1")
priors['addstepasym_2'] = bilby.core.prior.Uniform(0, 1, name="addstepasym_2")
for mix_model in mix_models:
mix_model.set_prior(priors)
for mix__model in mix_models:
print(mix_model.prior)
{'addstepasym_0': Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None), 'addstepasym_1': Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None), 'addstepasym_2': Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None), 'model1_0': Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None), 'model2_0': Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)}
{'addstepasym_0': Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None), 'addstepasym_1': Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None), 'addstepasym_2': Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None), 'model1_0': Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None), 'model2_0': Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)}
{'addstepasym_0': Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None), 'addstepasym_1': Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None), 'addstepasym_2': Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None), 'model1_0': Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None), 'model2_0': Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)}
2.5.3. 3. Train to find posterior#
g.shape
(10,)
#from Taweret.utils.utils import normed_mvn_loglike
kwargs_for_sampler = {'sampler':'ptemcee',
'ntemps':5,
'nwalkers':40,
'Tmax':100,
'burn_in_fixed_discard':500,
'nsamples':3000,
'threads':6,
'verbose':False}
#'safety':2,
#'autocorr_tol':5}
import os
import shutil
outdirs = ['outdir/mix_model_1', 'outdir/mix_model_2', 'outdir/mix_model_3']
labels = ['BMMC','BMMcor','BMMmean']
results = []
for i in range(0,3):
mix_model = mix_models[i]
label = labels[i]
outdir = outdirs[i]
if os.path.isdir(outdir):
print('removing '+outdir)
shutil.rmtree(outdir)
else:
print('file does not exist '+outdir)
result = mix_model.train(x_exp=g.reshape(-1,1), y_exp=exp_data[0].reshape(-1,1), y_err=exp_data[1].reshape(-1,1)
,kwargs_for_sampler=kwargs_for_sampler, label=label, outdir=outdir)
results.append(result)
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/likelihood.py:127: FutureWarning: Setting non-trivial parameters for <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>. This is deprecated behaviour and will be removed in Bilby version 3. See https://bilby-dev.github.io/bilby/parameters for more details.
warnings.warn(msg, FutureWarning)
19:35 bilby INFO : Running for label 'BMMC', output will be saved to 'outdir/mix_model_1'
file does not exist outdir/mix_model_1
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/sampler/ptemcee.py:134: FutureWarning: The ptemcee sampler interface in bilby is deprecated and will be removed in Bilby version 3. Please use the `ptemcee-bilby`sampler plugin instead: https://github.com/bilby-dev/ptemcee-bilby.
warnings.warn(msg, FutureWarning)
19:35 bilby INFO : Analysis priors:
19:35 bilby INFO : addstepasym_0=Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None)
19:35 bilby INFO : addstepasym_1=Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None)
19:35 bilby INFO : addstepasym_2=Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None)
19:35 bilby INFO : model1_0=Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None)
19:35 bilby INFO : model2_0=Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)
19:35 bilby INFO : Analysis likelihood class: <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>
19:35 bilby INFO : Analysis likelihood noise evidence: nan
19:35 bilby INFO : Single likelihood evaluation took 2.904e-04 s
19:35 bilby INFO : Using sampler Ptemcee with kwargs {'ntemps': 5, 'nwalkers': 40, 'Tmax': 100, 'betas': None, 'a': 2.0, 'adaptation_lag': 10000, 'adaptation_time': 100, 'random': None, 'adapt': False, 'swap_ratios': False}
19:35 bilby INFO : Global meta data was removed from the result object for compatibility. Use the `BILBY_INCLUDE_GLOBAL_METADATA` environment variable to include it. This behaviour will be removed in a future release. For more details see: https://bilby-dev.github.io/bilby/faq.html#global-meta-data
19:35 bilby INFO : Using convergence inputs: ConvergenceInputs(autocorr_c=5, autocorr_tol=50, autocorr_tau=1, gradient_tau=0.1, gradient_mean_log_posterior=0.1, Q_tol=1.02, safety=1, burn_in_nact=50, burn_in_fixed_discard=500, mean_logl_frac=0.01, thin_by_nact=0.5, nsamples=3000, ignore_keys_for_tau=None, min_tau=1, niterations_per_check=5)
19:35 bilby INFO : Generating pos0 samples
19:35 bilby INFO : Starting to sample
19:36 bilby INFO : Finished sampling
19:36 bilby INFO : Writing checkpoint and diagnostics
19:36 bilby INFO : Finished writing checkpoint
19:36 bilby INFO : Sampling time: 0:01:24.179991
19:36 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:36 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:36 bilby INFO : Summary of results:
nsamples: 3240
ln_noise_evidence: nan
ln_evidence: -8.957 +/- 2.525
ln_bayes_factor: nan +/- 2.525
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/likelihood.py:127: FutureWarning: Setting non-trivial parameters for <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>. This is deprecated behaviour and will be removed in Bilby version 3. See https://bilby-dev.github.io/bilby/parameters for more details.
warnings.warn(msg, FutureWarning)
19:36 bilby INFO : Running for label 'BMMcor', output will be saved to 'outdir/mix_model_2'
file does not exist outdir/mix_model_2
19:36 bilby INFO : Analysis priors:
19:36 bilby INFO : addstepasym_0=Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None)
19:36 bilby INFO : addstepasym_1=Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None)
19:36 bilby INFO : addstepasym_2=Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None)
19:36 bilby INFO : model1_0=Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None)
19:36 bilby INFO : model2_0=Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)
19:36 bilby INFO : Analysis likelihood class: <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>
19:36 bilby INFO : Analysis likelihood noise evidence: nan
19:36 bilby INFO : Single likelihood evaluation took 2.268e-04 s
19:36 bilby INFO : Using sampler Ptemcee with kwargs {'ntemps': 5, 'nwalkers': 40, 'Tmax': 100, 'betas': None, 'a': 2.0, 'adaptation_lag': 10000, 'adaptation_time': 100, 'random': None, 'adapt': False, 'swap_ratios': False}
19:36 bilby INFO : Global meta data was removed from the result object for compatibility. Use the `BILBY_INCLUDE_GLOBAL_METADATA` environment variable to include it. This behaviour will be removed in a future release. For more details see: https://bilby-dev.github.io/bilby/faq.html#global-meta-data
19:36 bilby INFO : Using convergence inputs: ConvergenceInputs(autocorr_c=5, autocorr_tol=50, autocorr_tau=1, gradient_tau=0.1, gradient_mean_log_posterior=0.1, Q_tol=1.02, safety=1, burn_in_nact=50, burn_in_fixed_discard=500, mean_logl_frac=0.01, thin_by_nact=0.5, nsamples=3000, ignore_keys_for_tau=None, min_tau=1, niterations_per_check=5)
19:36 bilby INFO : Generating pos0 samples
19:36 bilby INFO : Starting to sample
19:38 bilby INFO : Finished sampling
19:38 bilby INFO : Writing checkpoint and diagnostics
19:38 bilby INFO : Finished writing checkpoint
19:38 bilby INFO : Sampling time: 0:02:04.865254
19:38 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:38 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:38 bilby INFO : Summary of results:
nsamples: 3720
ln_noise_evidence: nan
ln_evidence: 7.103 +/- 5.062
ln_bayes_factor: nan +/- 5.062
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/likelihood.py:127: FutureWarning: Setting non-trivial parameters for <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>. This is deprecated behaviour and will be removed in Bilby version 3. See https://bilby-dev.github.io/bilby/parameters for more details.
warnings.warn(msg, FutureWarning)
19:38 bilby INFO : Running for label 'BMMmean', output will be saved to 'outdir/mix_model_3'
file does not exist outdir/mix_model_3
19:38 bilby INFO : Analysis priors:
19:38 bilby INFO : addstepasym_0=Uniform(minimum=0, maximum=9, name='addstepasym_0', latex_label='addstepasym_0', unit=None, boundary=None)
19:38 bilby INFO : addstepasym_1=Uniform(minimum=0, maximum=9, name='addstepasym_1', latex_label='addstepasym_1', unit=None, boundary=None)
19:38 bilby INFO : addstepasym_2=Uniform(minimum=0, maximum=1, name='addstepasym_2', latex_label='addstepasym_2', unit=None, boundary=None)
19:38 bilby INFO : model1_0=Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None)
19:38 bilby INFO : model2_0=Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)
19:38 bilby INFO : Analysis likelihood class: <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>
19:38 bilby INFO : Analysis likelihood noise evidence: nan
19:38 bilby INFO : Single likelihood evaluation took 1.889e-04 s
19:38 bilby INFO : Using sampler Ptemcee with kwargs {'ntemps': 5, 'nwalkers': 40, 'Tmax': 100, 'betas': None, 'a': 2.0, 'adaptation_lag': 10000, 'adaptation_time': 100, 'random': None, 'adapt': False, 'swap_ratios': False}
19:38 bilby INFO : Global meta data was removed from the result object for compatibility. Use the `BILBY_INCLUDE_GLOBAL_METADATA` environment variable to include it. This behaviour will be removed in a future release. For more details see: https://bilby-dev.github.io/bilby/faq.html#global-meta-data
19:38 bilby INFO : Using convergence inputs: ConvergenceInputs(autocorr_c=5, autocorr_tol=50, autocorr_tau=1, gradient_tau=0.1, gradient_mean_log_posterior=0.1, Q_tol=1.02, safety=1, burn_in_nact=50, burn_in_fixed_discard=500, mean_logl_frac=0.01, thin_by_nact=0.5, nsamples=3000, ignore_keys_for_tau=None, min_tau=1, niterations_per_check=5)
19:38 bilby INFO : Generating pos0 samples
19:38 bilby INFO : Starting to sample
19:40 bilby INFO : Finished sampling
19:40 bilby INFO : Writing checkpoint and diagnostics
19:40 bilby INFO : Finished writing checkpoint
19:40 bilby INFO : Sampling time: 0:01:27.096973
19:40 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:40 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:40 bilby INFO : Summary of results:
nsamples: 3040
ln_noise_evidence: nan
ln_evidence: -1.214 +/- 3.631
ln_bayes_factor: nan +/- 3.631
posteriors = [0,0,0]
for i in range(0,3):
result = results[i]
label = labels[i]
result = result.posterior.iloc[:,0:-2]
result['model'] = label
posteriors[i]=result
import pandas as pd
df = pd.concat(posteriors, ignore_index=True, sort=False)
df.head(-10)
| addstepasym_0 | addstepasym_1 | addstepasym_2 | model1_0 | model2_0 | model | |
|---|---|---|---|---|---|---|
| 0 | 2.486527 | 7.941589 | 0.662796 | 5.542920 | 0.976820 | BMMC |
| 1 | 3.097445 | 6.956633 | 0.789695 | 4.455517 | 0.881384 | BMMC |
| 2 | 3.430944 | 7.229666 | 0.863489 | 4.690683 | 1.299283 | BMMC |
| 3 | 3.461479 | 7.118759 | 0.855797 | 4.728741 | 1.325310 | BMMC |
| 4 | 3.912860 | 6.651543 | 0.870824 | 4.571047 | 1.443501 | BMMC |
| ... | ... | ... | ... | ... | ... | ... |
| 9985 | 3.848702 | 0.460433 | 0.992055 | 5.103604 | 1.501484 | BMMmean |
| 9986 | 3.884630 | 5.423056 | 0.990370 | 4.689796 | 1.283540 | BMMmean |
| 9987 | 3.363562 | 7.162613 | 0.996488 | 5.167660 | 1.053737 | BMMmean |
| 9988 | 3.394550 | 6.904732 | 0.991850 | 5.186315 | 1.081165 | BMMmean |
| 9989 | 3.580620 | 4.098574 | 0.985367 | 4.716994 | 1.617868 | BMMmean |
9990 rows × 6 columns
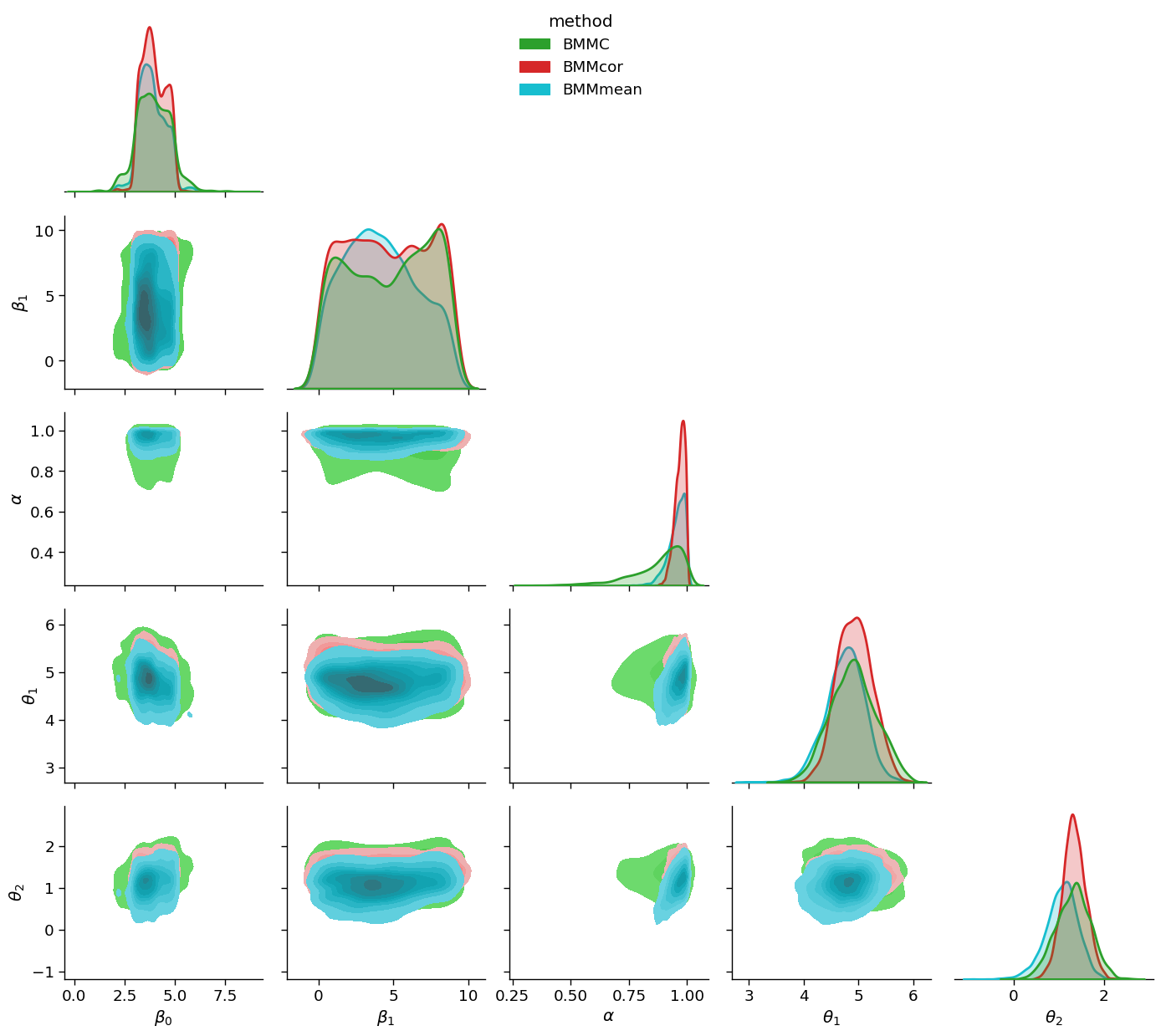
df_renamed=df.rename(columns={'addstepasym_0':r'$\beta_0$', 'addstepasym_1':r'$\beta_1$',
'addstepasym_2':r'$\alpha$', 'model1_0':r'$\theta_1$',
'model2_0':r'$\theta_2$', 'model':'method'})
#g.savefig('temp_save')
import seaborn as sns
sns.set_context('paper', font_scale=1.5)
gg = sns.PairGrid(df_renamed, hue="method", diag_sharey=False, hue_kws={'alpha':0.5}, corner=True,
palette={'BMMC':sns.color_palette()[2],'BMMcor':sns.color_palette()[3], 'BMMmean':sns.color_palette()[-1]})
gg.map_lower(sns.kdeplot, fill=True)
gg.map_diag(sns.kdeplot, linewidth=2, fill=True)
gg.add_legend(loc='upper center')
plt.tight_layout()
plt.savefig('comparative_posterior', dpi=100)
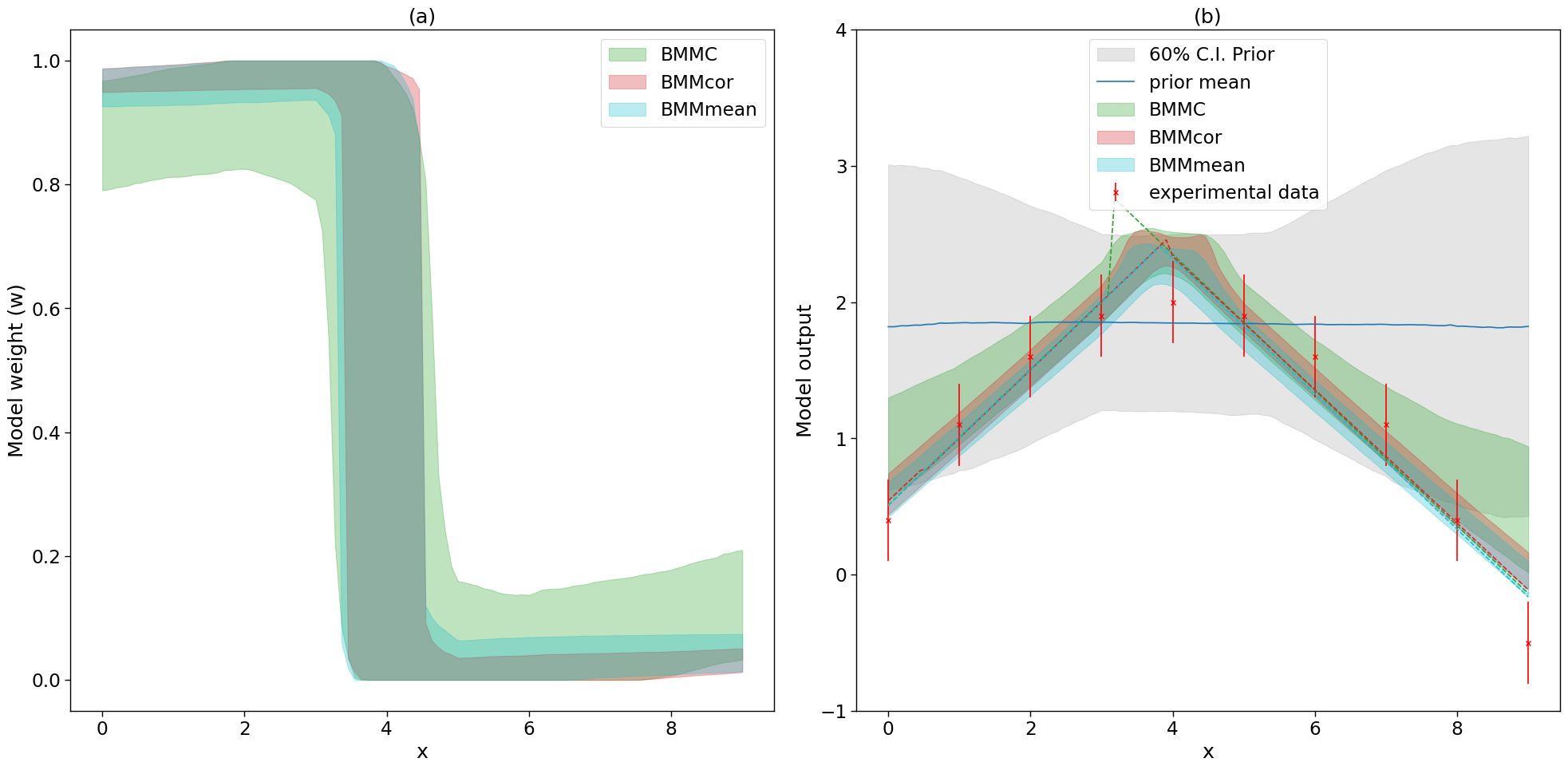
2.5.3.1. 4. Predictions#
sns.set_context('paper', font_scale=1.9)
fig, axs = plt.subplots(1,2,figsize=(20,10))
ax, ax2 = axs.flatten()
#fig2, ax2 = plt.subplots(figsize=(10,10))
colors = {'BMMC':sns.color_palette()[2],'BMMcor':sns.color_palette()[3], 'BMMmean':sns.color_palette()[-1]}
for i, mix_model in enumerate(mix_models):
_,mean_prior,CI_prior, _ = mix_model.prior_predict(plot_g, CI=[5,20,80,95])
_,mean,CI, _ = mix_model.predict(plot_g, CI=[5,20,80,95])
per5, per20, per80, per95 = CI
prior5, prior20, prior80, prior95 = CI_prior
# Map value prediction for the step mixing function parameter
model_params = [np.array(mix_model.map[3]), np.array(mix_model.map[4])]
map_prediction = mix_model.evaluate(mix_model.map[0:3], plot_g, model_params=model_params)
print(mix_model.map)
_,_,CI_weights,_=mix_model.predict_weights(plot_g, CI=[5,20, 80, 95])
perw_5, perw_20, perw_80, perw_95 = CI_weights
#ax.fill_between(plot_g,perw_5,perw_95,color=colors[labels[i]], alpha=0.2, label='90% C.I.')
ax.fill_between(plot_g,perw_20,perw_80, color=colors[labels[i]], alpha=0.3, label=labels[i])
if i==0:
ax2.fill_between(plot_g,prior20.flatten(),prior80.flatten(),color=sns.color_palette()[7], alpha=0.2, label='60% C.I. Prior')
ax2.errorbar(g,exp_data[0],yerr=exp_data[1], marker='x', label='experimental data', color='red', fmt='.')
ax2.plot(plot_g, mean_prior.flatten(), label='prior mean')
#ax2.plot(plot_g, mean.flatten(), label=labels[i])
#ax2.fill_between(plot_g,per5.flatten(),per95.flatten(),color=sns.color_palette()[4], alpha=0.2, label='90% C.I.')
ax2.fill_between(plot_g,per20.flatten(),per80.flatten(), color=colors[labels[i]], alpha=0.3, label=labels[i])
ax2.plot(plot_g, map_prediction.flatten(), color=colors[labels[i]], linestyle='dashed')
ax.legend()
ax.set_xlabel('x')
ax.set_ylabel('Model weight (w)')
ax2.set_ybound(-1,4)
ax2.legend(loc='upper center')
ax2.set_xlabel('x')
ax2.set_ylabel('Model output')
ax.set_title('(a)')
ax2.set_title('(b)')
plt.tight_layout()
fig.savefig('comparative_posterior_prditcions', dpi=100)
#fig2.savefig('comparative_posterior_predict', dpi=100)
[3.11361453 7.44964417 0.99807884 5.00308475 1.30365511]
/tmp/ipykernel_4738/1972745161.py:26: UserWarning: marker is redundantly defined by the 'marker' keyword argument and the fmt string "." (-> marker='.'). The keyword argument will take precedence.
ax2.errorbar(g,exp_data[0],yerr=exp_data[1], marker='x', label='experimental data', color='red', fmt='.')
[3.99225858 0.51047784 0.98947215 5.00438811 1.26912869]
[3.83041157 4.92808133 0.99695917 4.99453917 1.24114518]