2.4. Linear Bivariate BMM with calibration of Coleman toy models.#
The best way to learn Taweret is to use it. You can run, modify and experiment with this notebook on GitHub Codespaces.
The models can be found in Coleman Thesis : https://go.exlibris.link/3fVZCfhl
This notebook shows how to use the Bayesian model mixing package Taweret for a toy problem.
Author : Dan Liyanage
Date : 11/13/2022
import sys
import os
# You will have to change the following imports depending on where you have
# the packages installed
# ! pip install Taweret # if using Colab, uncomment to install
cwd = os.getcwd()
# Get the first part of this path and append to the sys.path
tw_path = cwd.split("Taweret/")[0] + "Taweret"
sys.path.append(tw_path)
# For plotting
import matplotlib.pyplot as plt
! pip install seaborn # comment if installed
! pip install ptemcee # comment if installed
import seaborn as sns
sns.set_context('poster')
# To define priors. (uncoment if not using default priors)
# ! pip install bilby # uncomment this line if bilby is not already installed
# import bilby
# For other operations
import numpy as np
Requirement already satisfied: seaborn in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (0.13.2)
Requirement already satisfied: numpy!=1.24.0,>=1.20 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (2.4.6)
Requirement already satisfied: pandas>=1.2 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (3.0.3)
Requirement already satisfied: matplotlib!=3.6.1,>=3.4 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from seaborn) (3.10.9)
Requirement already satisfied: contourpy>=1.0.1 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (1.3.3)
Requirement already satisfied: cycler>=0.10 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (0.12.1)
Requirement already satisfied: fonttools>=4.22.0 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (4.63.0)
Requirement already satisfied: kiwisolver>=1.3.1 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (1.5.0)
Requirement already satisfied: packaging>=20.0 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (26.2)
Requirement already satisfied: pillow>=8 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (12.2.0)
Requirement already satisfied: pyparsing>=3 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (3.3.2)
Requirement already satisfied: python-dateutil>=2.7 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from matplotlib!=3.6.1,>=3.4->seaborn) (2.9.0.post0)
Requirement already satisfied: six>=1.5 in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from python-dateutil>=2.7->matplotlib!=3.6.1,>=3.4->seaborn) (1.17.0)
Requirement already satisfied: ptemcee in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (1.0.0)
Requirement already satisfied: numpy in /home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages (from ptemcee) (2.4.6)
# Import models with a predict method
from Taweret.models import coleman_models as toy_models
m1 = toy_models.coleman_model_1()
m2 = toy_models.coleman_model_2()
truth = toy_models.coleman_truth()
g = np.linspace(-1,9,10)
plot_g = np.linspace(-1,9,100)
true_output = truth.evaluate(plot_g)
exp_data = truth.evaluate(g)
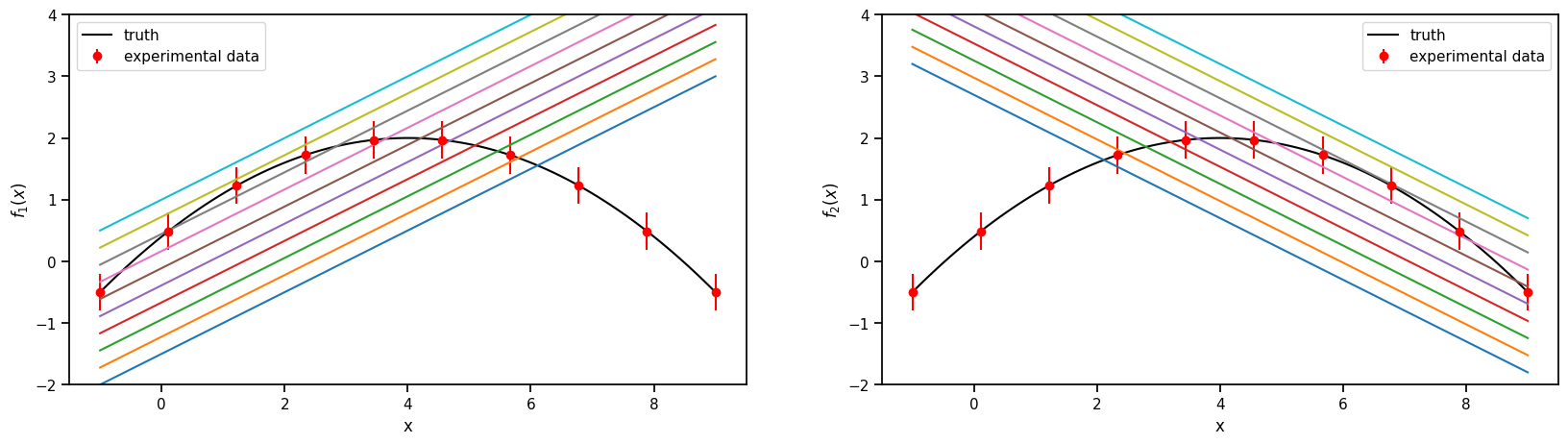
2.4.1. 1. The models and the experimental data.#
Truth
\(f(x) = 2-0.1(x-4)^2\), where \(x \in [-1, 9]\)
Model 1
\(f_1(x,\theta)= 0.5(x+\theta)-2\) , where \(\theta \in [1, 6]\)
Model 2
\(f_2(x,\theta)= -0.5(x-\theta) + 3.7\) , where \(\theta \in [-2, 3]\)
Experimental data
sampled from the Truth with a fixed standard deviation of 0.3
sns.set_context('notebook')
fig, axs = plt.subplots(1,2,figsize=(20,5))
prior_ranges = [(1,6), (-2,3)]
for i in range(0,2):
ax = axs.flatten()[i]
ax.plot(plot_g, true_output[0], label='truth', color='black')
ax.errorbar(g,exp_data[0],exp_data[1], fmt='o', label='experimental data', color='r')
ax.legend()
ax.set_ylim(-2,4)
for value in np.linspace(*prior_ranges[i],10):
if i==0:
predict_1 = m1.evaluate(plot_g, value)
ax.plot(plot_g, predict_1[0])
ax.set_ylabel(r'$f_1(x)$')
if i==1:
predict_2 = m2.evaluate(plot_g, value)
ax.plot(plot_g, predict_2[0])
ax.set_ylabel(r'$f_2(x)$')
ax.set_xlabel('x')
2.4.2. 2. Choose a Mixing method#
# Mixing method
from Taweret.mix.bivariate_linear import BivariateLinear as BL
models= {'model1':m1,'model2':m2}
mix_model = BL(models_dic=models, method='cdf', nargs_model_dic={'model1':1, 'model2':1})
## uncoment to change the prior from the default
import bilby
priors = bilby.core.prior.PriorDict()
priors['cdf_0'] = bilby.core.prior.Normal(0, 1, name="cdf_0")
priors['cdf_1'] = bilby.core.prior.Normal(0, 1, name="cdf_1")
mix_model.set_prior(priors)
{'cdf_0': Normal(mu=0, sigma=1, name='cdf_0', latex_label='cdf_0', unit=None, boundary=None),
'cdf_1': Normal(mu=0, sigma=1, name='cdf_1', latex_label='cdf_1', unit=None, boundary=None),
'model1_0': Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None),
'model2_0': Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)}
mix_model.prior
{'cdf_0': Normal(mu=0, sigma=1, name='cdf_0', latex_label='cdf_0', unit=None, boundary=None),
'cdf_1': Normal(mu=0, sigma=1, name='cdf_1', latex_label='cdf_1', unit=None, boundary=None),
'model1_0': Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None),
'model2_0': Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)}
2.4.3. 3. Train to find posterior#
kwargs_for_sampler = {'sampler': 'ptemcee',
'ntemps': 5,
'nwalkers': 50,
'Tmax': 100,
'burn_in_fixed_discard': 50,
'nsamples': 2000,
'threads': 6,
'verbose': False}
result = mix_model.train(x_exp=g, y_exp=exp_data[0], y_err=exp_data[1],
label='cdf_mix',
outdir='outdir/coleman',
kwargs_for_sampler=kwargs_for_sampler,
load_previous=False)
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/likelihood.py:127: FutureWarning: Setting non-trivial parameters for <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>. This is deprecated behaviour and will be removed in Bilby version 3. See https://bilby-dev.github.io/bilby/parameters for more details.
warnings.warn(msg, FutureWarning)
19:29 bilby INFO : Running for label 'cdf_mix', output will be saved to 'outdir/coleman'
/home/runner/work/Taweret/Taweret/.tox/book/lib/python3.13/site-packages/bilby/core/sampler/ptemcee.py:134: FutureWarning: The ptemcee sampler interface in bilby is deprecated and will be removed in Bilby version 3. Please use the `ptemcee-bilby`sampler plugin instead: https://github.com/bilby-dev/ptemcee-bilby.
warnings.warn(msg, FutureWarning)
19:29 bilby INFO : Analysis priors:
19:29 bilby INFO : cdf_0=Normal(mu=0, sigma=1, name='cdf_0', latex_label='cdf_0', unit=None, boundary=None)
19:29 bilby INFO : cdf_1=Normal(mu=0, sigma=1, name='cdf_1', latex_label='cdf_1', unit=None, boundary=None)
19:29 bilby INFO : model1_0=Uniform(minimum=1, maximum=6, name='model1_0', latex_label='model1_0', unit=None, boundary=None)
19:29 bilby INFO : model2_0=Uniform(minimum=-2, maximum=3, name='model2_0', latex_label='model2_0', unit=None, boundary=None)
19:29 bilby INFO : Analysis likelihood class: <class 'Taweret.sampler.likelihood_wrappers.likelihood_wrapper_for_bilby'>
19:29 bilby INFO : Analysis likelihood noise evidence: nan
19:29 bilby INFO : Single likelihood evaluation took 3.318e-04 s
19:29 bilby INFO : Using sampler Ptemcee with kwargs {'ntemps': 5, 'nwalkers': 50, 'Tmax': 100, 'betas': None, 'a': 2.0, 'adaptation_lag': 10000, 'adaptation_time': 100, 'random': None, 'adapt': False, 'swap_ratios': False}
19:29 bilby INFO : Global meta data was removed from the result object for compatibility. Use the `BILBY_INCLUDE_GLOBAL_METADATA` environment variable to include it. This behaviour will be removed in a future release. For more details see: https://bilby-dev.github.io/bilby/faq.html#global-meta-data
19:29 bilby INFO : Using convergence inputs: ConvergenceInputs(autocorr_c=5, autocorr_tol=50, autocorr_tau=1, gradient_tau=0.1, gradient_mean_log_posterior=0.1, Q_tol=1.02, safety=1, burn_in_nact=50, burn_in_fixed_discard=50, mean_logl_frac=0.01, thin_by_nact=0.5, nsamples=2000, ignore_keys_for_tau=None, min_tau=1, niterations_per_check=5)
19:29 bilby INFO : Generating pos0 samples
19:29 bilby INFO : Starting to sample
19:30 bilby INFO : Finished sampling
19:30 bilby INFO : Writing checkpoint and diagnostics
19:30 bilby INFO : Finished writing checkpoint
19:30 bilby INFO : Sampling time: 0:01:26.046969
19:30 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:30 bilby WARNING : Result.save_to_file called with extension=True. This will default to json, and ignore the extension from the filename. This behaviour is deprecated and will be removed.
19:30 bilby INFO : Summary of results:
nsamples: 7450
ln_noise_evidence: nan
ln_evidence: -10.410 +/- 2.376
ln_bayes_factor: nan +/- 2.376
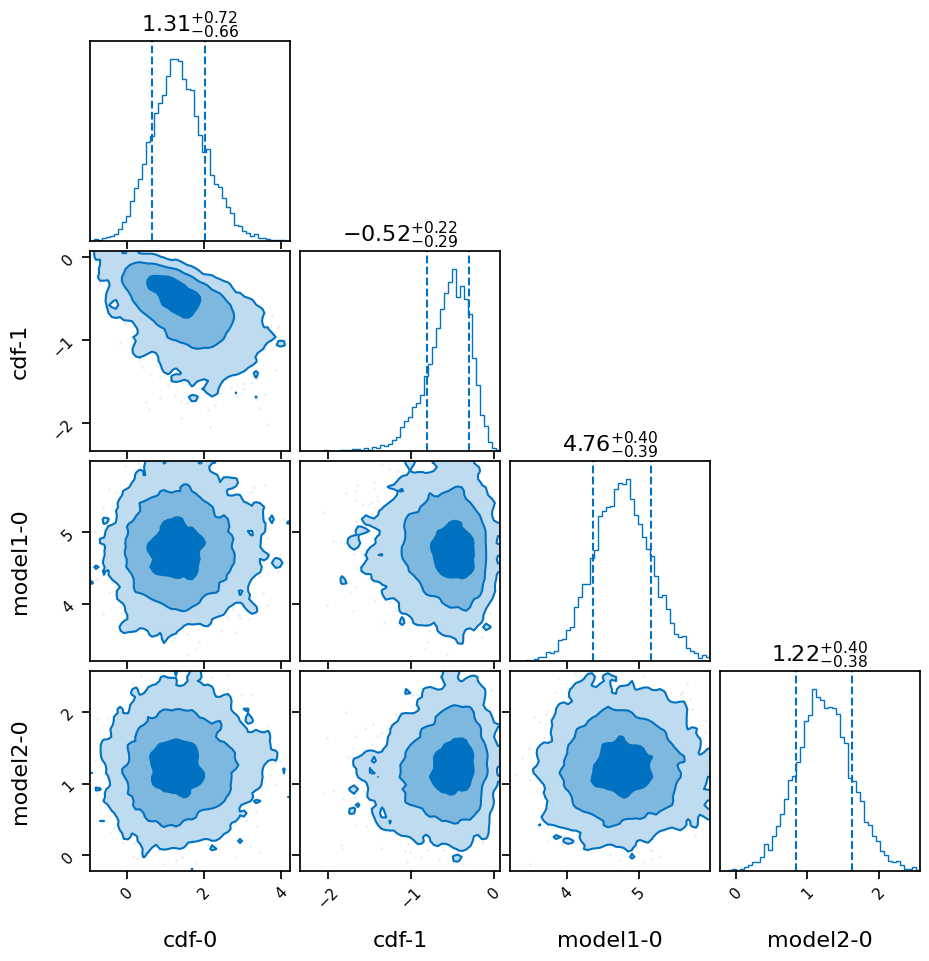
result.plot_corner()
2.4.3.1. 4. Predictions#
_,mean_prior,CI_prior, _ = mix_model.prior_predict(plot_g, CI=[5,20,80,95])
_,mean,CI, _ = mix_model.predict(plot_g, CI=[5,20,80,95])
per5, per20, per80, per95 = CI
prior5, prior20, prior80, prior95 = CI_prior
mix_model.map
array([ 3.95616515, -1.04956112, 4.68024204, 1.30910379])
# Map value prediction for the step mixing function parameter
model_params = [np.array(mix_model.map[2]), np.array(mix_model.map[3])]
map_prediction = mix_model.evaluate(mix_model.map[0:2], plot_g, model_params=model_params)
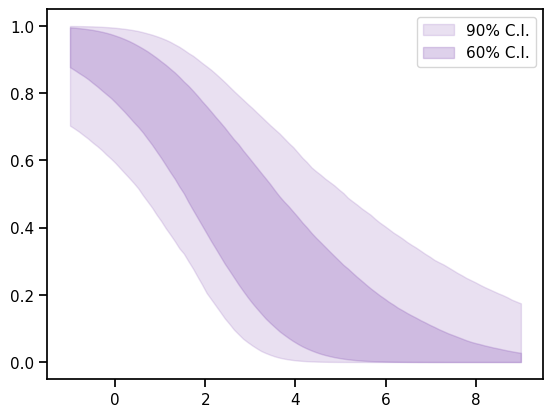
_,_,CI_weights,_=mix_model.predict_weights(plot_g, CI=[5,20, 80, 95])
perw_5, perw_20, perw_80, perw_95 = CI_weights
%matplotlib inline
fig, ax = plt.subplots()
ax.fill_between(plot_g,perw_5,perw_95,color=sns.color_palette()[4], alpha=0.2, label='90% C.I.')
ax.fill_between(plot_g,perw_20,perw_80, color=sns.color_palette()[4], alpha=0.3, label='60% C.I.')
ax.legend()
#ax.plot(plot_g, true_output[0], label='truth')
#ax.set_ylim(1.2,3.2)
<matplotlib.legend.Legend at 0x7f8615279950>
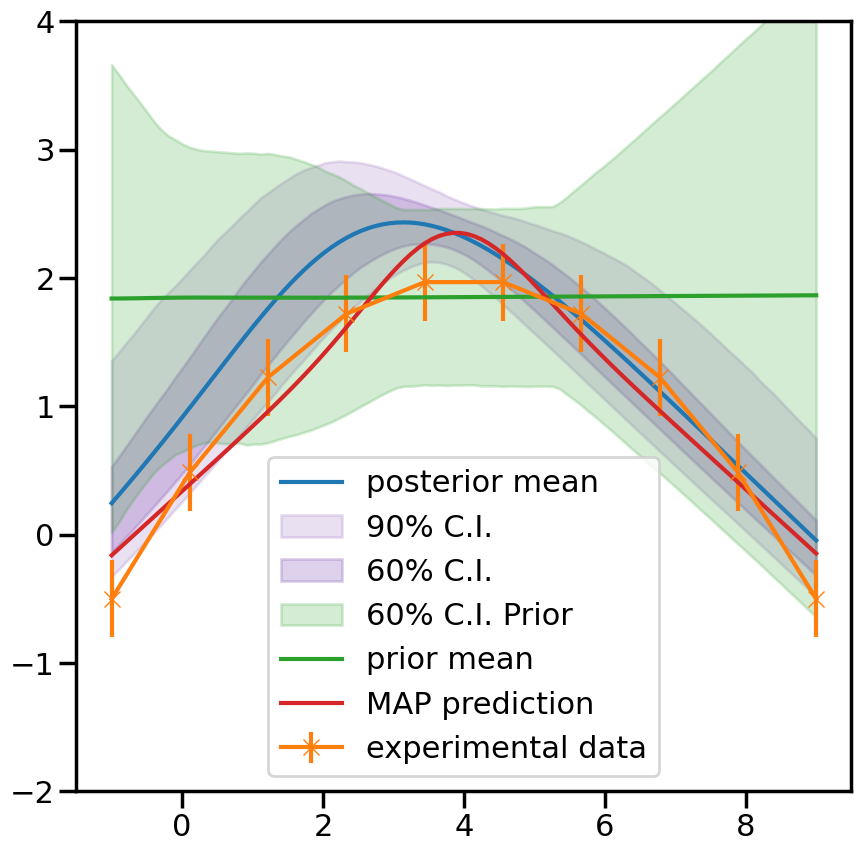
%matplotlib inline
sns.set_context('poster')
fig, ax = plt.subplots(figsize=(10,10))
ax.plot(plot_g, mean.flatten(), label='posterior mean')
ax.fill_between(plot_g,per5.flatten(),per95.flatten(),color=sns.color_palette()[4], alpha=0.2, label='90% C.I.')
ax.fill_between(plot_g,per20.flatten(),per80.flatten(), color=sns.color_palette()[4], alpha=0.3, label='60% C.I.')
ax.fill_between(plot_g,prior20.flatten(),prior80.flatten(),color=sns.color_palette()[2], alpha=0.2, label='60% C.I. Prior')
ax.errorbar(g,exp_data[0],yerr=exp_data[1], marker='x', label='experimental data')
ax.plot(plot_g, mean_prior.flatten(), label='prior mean')
ax.plot(plot_g, map_prediction.flatten(), label='MAP prediction')
ax.set_ybound(-2,4)
ax.legend()
<matplotlib.legend.Legend at 0x7f8615541bd0>